MAGEMin_C.jl: Fractional crystallization
This tutorial demonstrates how to perform fractional crystallization using MAGEMin_C.jl, from a hydrous basalt starting composition along a cooling and decompression path, with visualization of melt composition and density evolution.
Note
- This tutorial is not optimized for performance, but is provided in the hope it can be useful to present
MAGEMin_Cfunctionality.
Fractional crystallization
The following example shows how to perform fractional crystallization using a hydrous basalt magma as a starting composition. The results are displayed using PlotlyJS. This example is provided in the hope it may be useful for learning how to use MAGEMin_C for more advanced applications.
Note
Note that if we wanted to use a buffer, we would need to initialize MAGEMin as in example 4. Because during fractional crystallization the bulk-rock composition is updated at every step, we would need to increase the oxygen-content (O) of the new bulk-rock
using MAGEMin_C
using PlotlyJS
# number of computational steps
nsteps = 64
# Starting/ending Temperature [°C]
T = range(1200.0,600.0,nsteps)
# Starting/ending Pressure [kbar]
P = range(3.0,0.1,nsteps)
# Starting composition [mol fraction], here we used an hydrous basalt; composition taken from Blatter et al., 2013 (01SB-872, Table 1), with added O and water saturated
oxides = ["SiO2"; "Al2O3"; "CaO"; "MgO"; "FeO"; "K2O"; "Na2O"; "TiO2"; "O"; "Cr2O3"; "H2O"]
bulk_0 = [38.448328757254195, 7.718376151972274, 8.254653357127351, 9.95911842561036, 5.97899305676308, 0.24079752710315697, 2.2556006776515964, 0.7244006013202644, 0.7233140004182841, 0.0, 12.696417444779453];
# Define bulk-rock composition unit
sys_in = "mol"
# Choose database
data = Initialize_MAGEMin("ig", verbose=false);
# allocate storage space
Out_XY = Vector{out_struct}(undef,nsteps)
melt_F = 1.0
bulk = copy(bulk_0)
np = 0
while melt_F > 0.0
np +=1
out = single_point_minimization(P[np], T[np], data, X=bulk, Xoxides=oxides, sys_in=sys_in)
Out_XY[np] = deepcopy(out)
# retrieve melt composition to use as starting composition for next iteration
melt_F = out.frac_M
bulk .= out.bulk_M
print("#$np P: $(round(P[np],digits=3)), T: $(round(T[np],digits=3))\n")
print(" ---------------------\n")
print(" melt_F: $(round(melt_F, digits=3))\n melt_composition: $(round.(bulk ,digits=3))\n\n")
end
ndata = np -1 # last point has melt fraction = 0
x = Vector{String}(undef,ndata)
melt_SiO2_anhydrous = Vector{Float64}(undef,ndata)
melt_FeO_anhydrous = Vector{Float64}(undef,ndata)
melt_H2O = Vector{Float64}(undef,ndata)
fluid_frac = Vector{Float64}(undef,ndata)
melt_density = Vector{Float64}(undef,ndata)
residual_density = Vector{Float64}(undef,ndata)
system_density = Vector{Float64}(undef,ndata)
for i=1:ndata
x[i] = "[$(round(P[i],digits=3)), $(round(T[i],digits=3))]"
melt_SiO2_anhydrous[i] = Out_XY[i].bulk_M[1] / (sum(Out_XY[i].bulk_M[1:end-1])) * 100.0
melt_FeO_anhydrous[i] = Out_XY[i].bulk_M[5] / (sum(Out_XY[i].bulk_M[1:end-1])) * 100.0
melt_H2O[i] = Out_XY[i].bulk_M[end] *100
fluid_frac[i] = Out_XY[i].frac_F*100
melt_density[i] = Out_XY[i].rho_M
residual_density[i] = Out_XY[i].rho_S
system_density[i] = Out_XY[i].rho
end
# section to plot composition evolution
trace1 = scatter( x = x,
y = melt_SiO2_anhydrous,
name = "Anyhdrous SiO₂ [mol%]",
line = attr( color = "firebrick",
width = 2) )
trace2 = scatter( x = x,
y = melt_FeO_anhydrous,
name = "Anyhdrous FeO [mol%]",
line = attr( color = "royalblue",
width = 2) )
trace3 = scatter( x = x,
y = melt_H2O,
name = "H₂O [mol%]",
line = attr( color = "cornflowerblue",
width = 2) )
trace4 = scatter( x = x,
y = fluid_frac,
name = "fluid [mol%]",
line = attr( color = "black",
width = 2) )
layout = Layout( title = "Melt composition",
xaxis_title = "PT [kbar, °C]",
yaxis_title = "Oxide [mol%]")
plot([trace1,trace2,trace3,trace4], layout)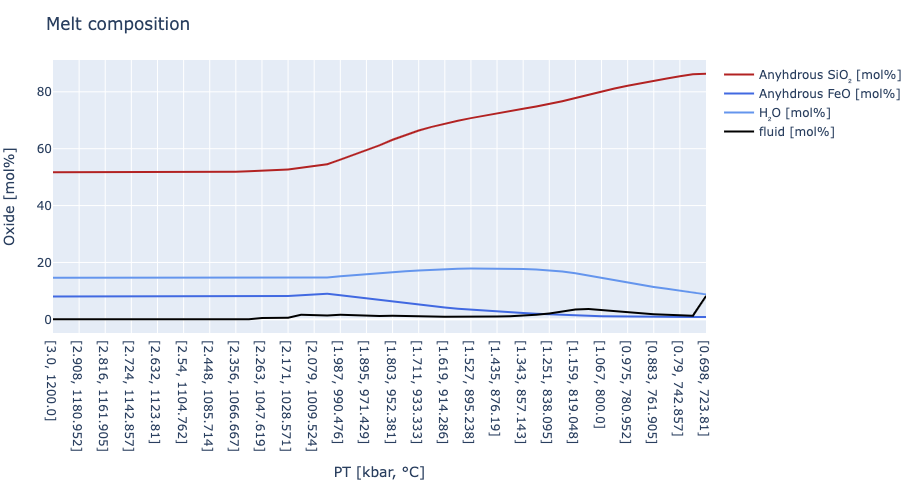
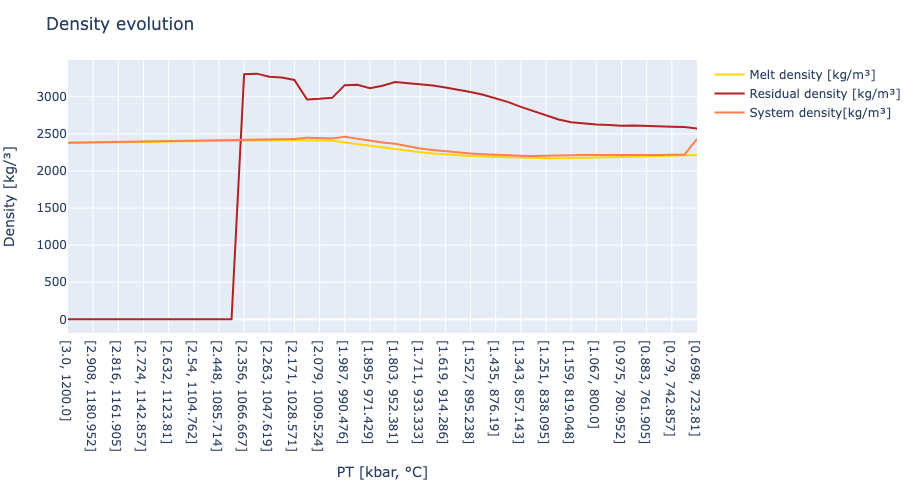
# section to plot density evolution
trace1 = scatter( x = x,
y = melt_density,
name = "Melt density [kg/m³]",
line = attr( color = "gold",
width = 2) )
trace2 = scatter( x = x,
y = residual_density,
name = "Residual density [kg/m³]",
line = attr( color = "firebrick",
width = 2) )
trace3 = scatter( x = x,
y = system_density,
name = "System density[kg/m³]",
line = attr( color = "coral",
width = 2) )
layout = Layout( title = "Density evolution",
xaxis_title = "PT [kbar, °C]",
yaxis_title = "Density [kg/³]")
plot([trace1,trace2,trace3], layout)